You are here
Software
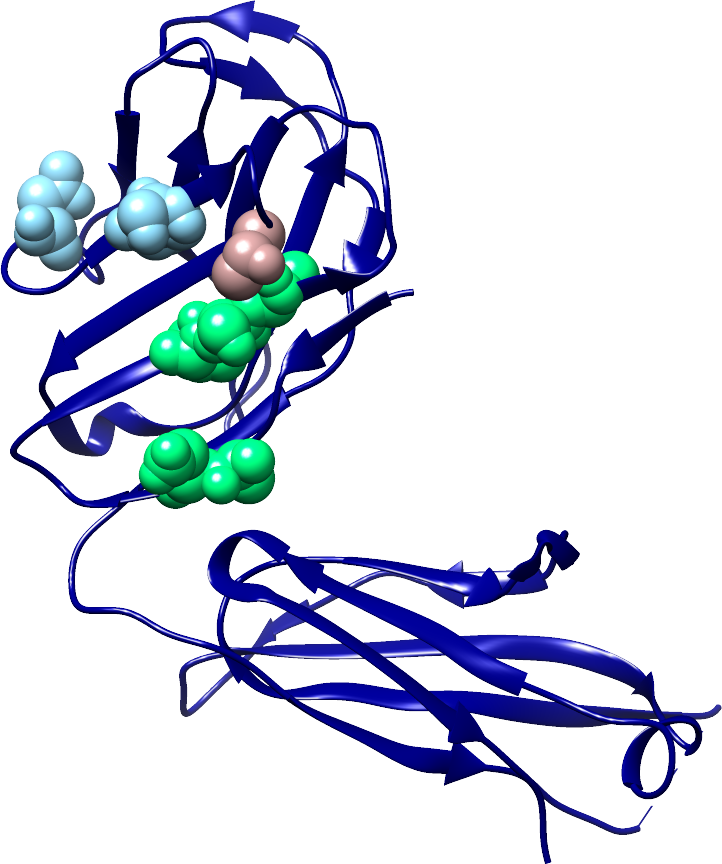 Co-evolution analysis of the envelope glycoproteins E1 and E2 of the Hepatitis C Virus (HCV) |
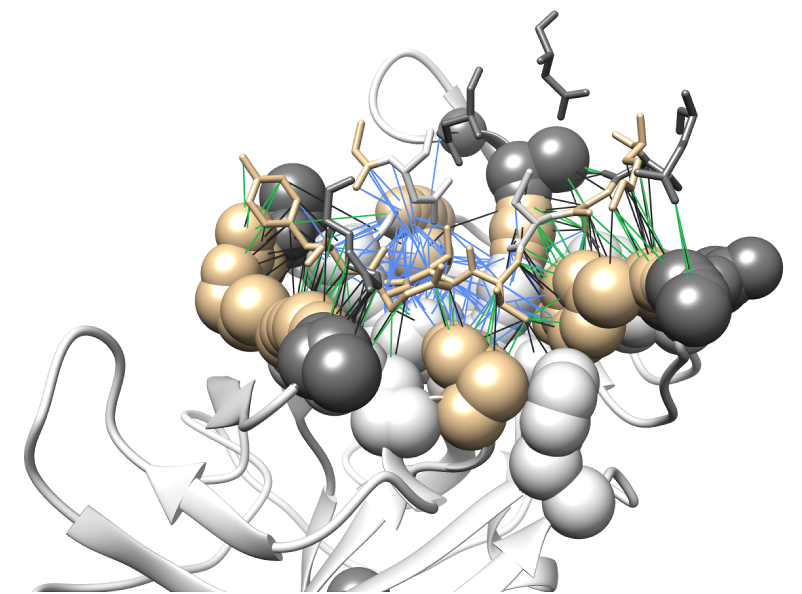 An empirical scoring function that estimates the binding affinity between two proteins, given the 3D structure of their complex. |
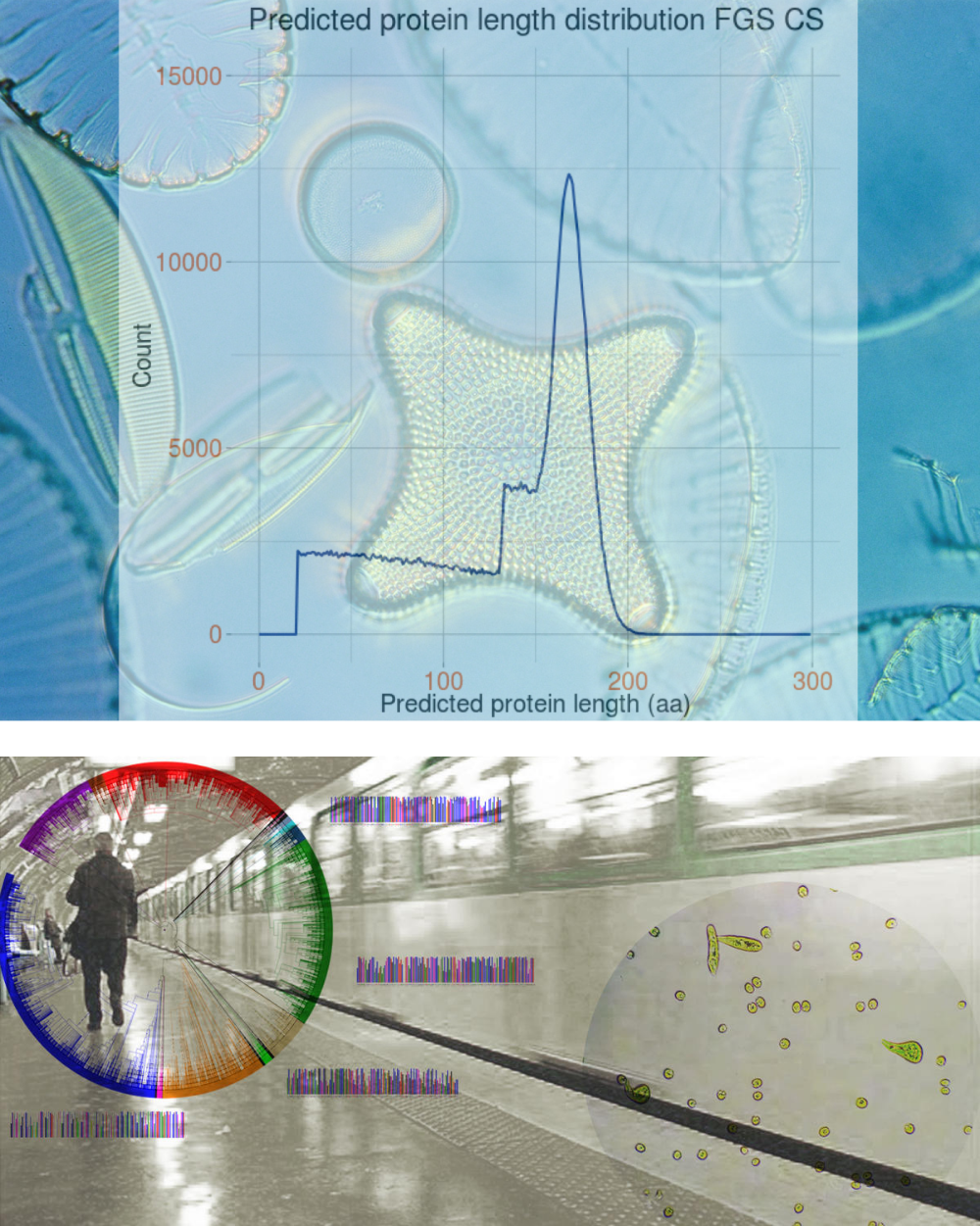 A novel profile-based domain annotation pipeline based on the multi-source domain annotation strategy. |
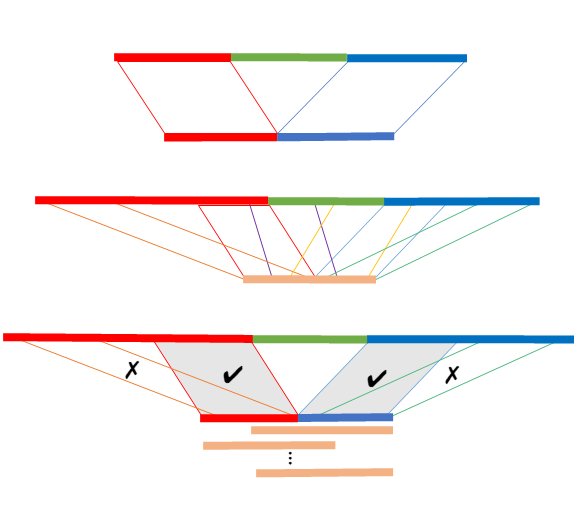 A split-alignment tool to infer structural variants (currently implemented for deletions only) from large short-read datasets. |
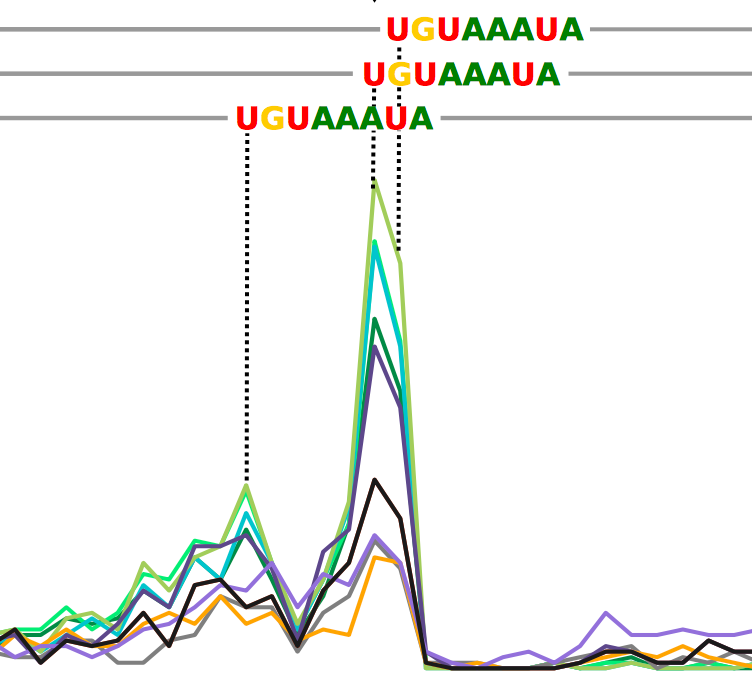 Capturing protein-RNA interaction footprints from single-nucleotide CLIP-seq data |




